Welcome to the TIGA Center
Tissue Imaging and Analysis Center
The TIGA Center is a cooperative project which started in 2007 at the University Heidelberg with the goal of establishing a bioinformatics platform dedicated to the quantitative analysis and modeling of tissues. A strong emphasis is placed on clinically relevant research projects.
At the heart of the TIGA’s technology platform are automated microscopic scanners for whole slide imaging of glass slides. By integrating such imaging systems in a technical pipeline ranging from organotypic in vitro cell cultures to computational tissue modeling the TIGA generates a wealth of yet unexploited clinically highly relevant tissue data.
We are able to process:
- tissue samples
- unstained histological slides
- stained histological slides
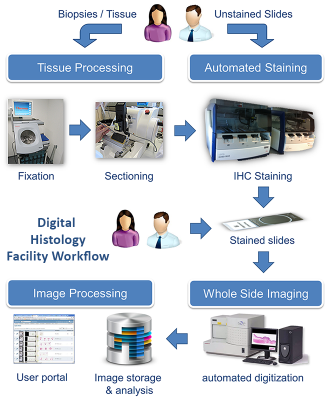
Tissue samples will be fixed, embedded and cut with the in-house histological equipment and prepared for automated staining. Tissue sections are stained with the Leica-Bond II Max staining machine.
In the following step the stained slides are digitized with one of the two Hamamatsu NanoZoomer Digital Pathology (NDP) systems (scanning a 15 mm x 15 mm region in 20x magnification). The specimen can be scanned with either 20x or 40x magnification, yielding a high spatial resolution of 0.46 µm/pixel and 0.23 µm/pixel respectively. The NDP systems detectors allow brightfield and fluorescence imaging of fixed tissue samples.
After scanning, the high resolution-images with a total file size of approximately 2 GB (uncompressed) per each single slide are presented on a dedicated whole slide image webserver (WSI database). We provide specific image processing solutions allowing to access WSI slide collections to perform standard image analysis or custom algorithms on the digitized slides.